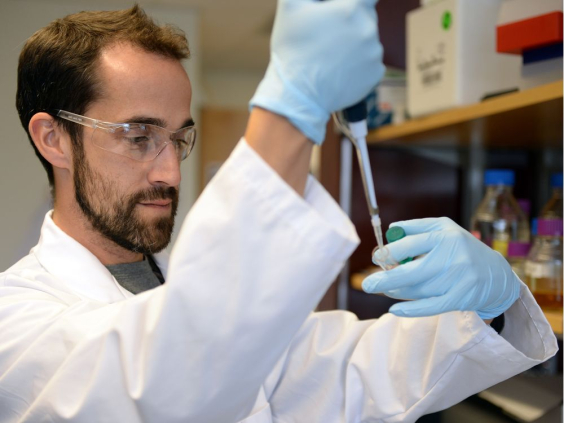
Welcome to IMSS
The Institute for Microbial Systems and Society (IMSS) unites functional microbial genetics research within the Faculty of Science and provides a platform to increase research activities and collaborations for research impact. The IMSS provides access to genomic technologies combined with the exploration of new ways to integrate genomic tools to collaborative research programs. The IMSS integrates current and emerging sequencing technologies to develop comprehensive approaches for studying the genetic makeup, gene expression, and gene function in microbial organisms and within microbial communities.
The IMSS has trained a large cohort of students and PDFs since its creation in 2016 by Dr. Andrew Cameron and Dr. Christopher Yost at the University of Regina. These include trainees that receive prestigious national scholarships, including NSERC Postdoctoral Fellowships, NSERC Canadian Graduate Scholarships, NSERC Undergraduate Student Research Awards, and provincial Postdoctoral Fellowships.
News
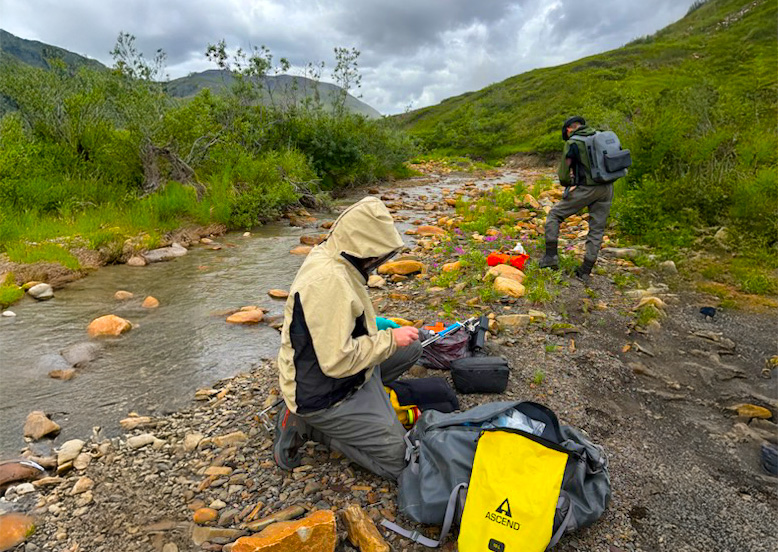

Tracking Invisible Threats: How Environmental DNA Could Transform Disease Surveillance on the Prairies
“Understanding how pathogens travel through these interconnected systems has long been a challenge for scientists and public health officials… Launched in late 2025, The Environmental DNA Surveillance Initiative (or eDNA Surveillance Project) seeks to develop practical tools that can monitor pathogens across agricultural systems, communities, and natural landscapes. The $1.83 million project with $475,000 in supports from Genome Canada, is co-led by Dr. Andrew Cameron, a microbial geneticist at the University of Regina, and Dr. Tony Ruzzini, Associate Professor at the Department of Veterinary Microbiology at the University of Saskatchewan.” Read the full article here.

IMSS member Rafael Grajales won People’s Choice and 3rd place at the U of R 3 Minute Thesis (3MT) competition!
Rafa presented his research on using microbial biocontrols to prevent root rot in pulse crops caused by the water mould Aphanomyces.
3MT is an annual competition where students present their research in under three minutes with only one static slide. First – third place winners are determined by a panel of judges from the community while People’s Choice is voted by the audience with paper ballots.
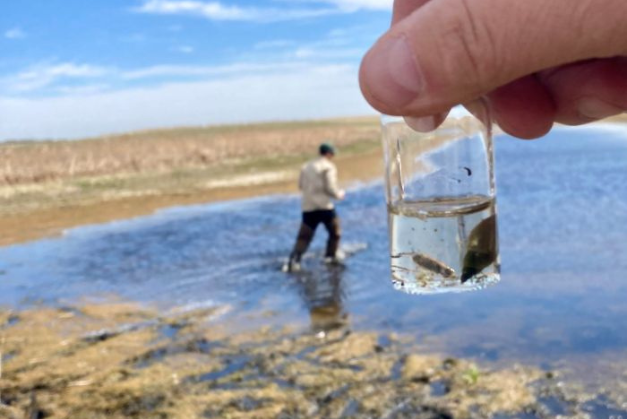
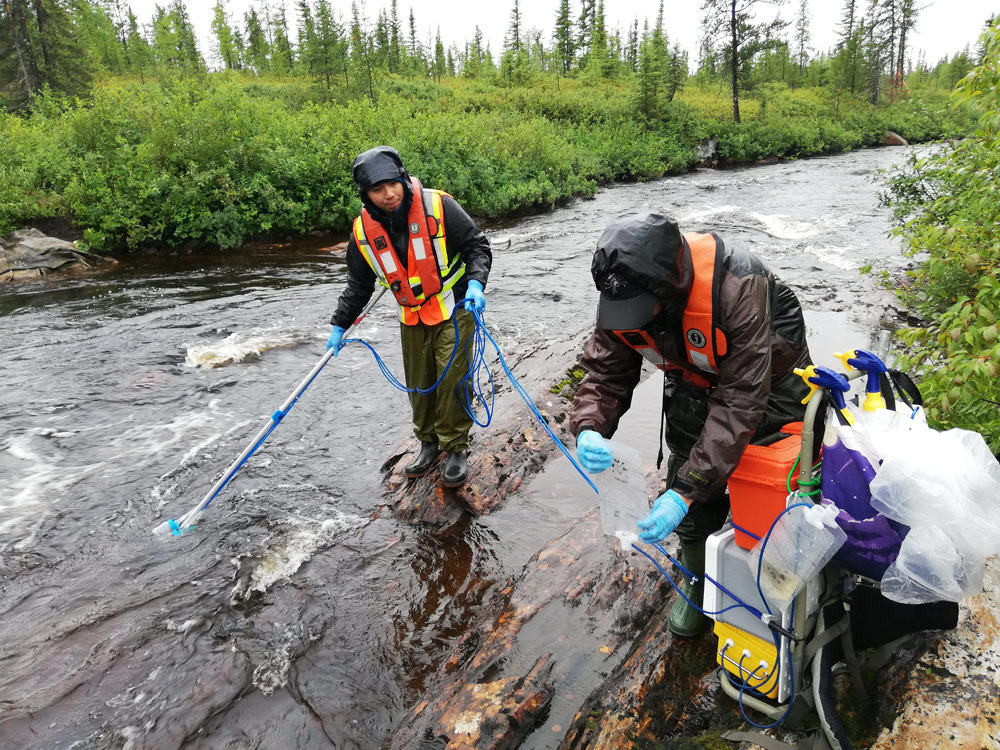



Genomic Innovations: eDNA & OneHealth Surveillance for a Safer Canada
Genome Canada is advancing innovative genomics tools that use environmental DNA (eDNA) and targeted metagenomics to detect biodiversity changes, pathogens, and antibiotic resistance from samples like water, soil, and wastewater. These non-invasive, highly sensitive methods will support communities – including northern, rural, remote, and Indigenous regions – in monitoring health threats across human, animal, and environmental interfaces, strengthening Canada’s capacity for rapid response and ecosystem protection.
“The “One Health”project, led by Drs. Andrew Cameron (University of Regina) and Tony Ruzzuini (University of Saskatchewan), will detect the presence of human and livestock pathogens, while monitoring the presence of AMR in the environment. The team will address technical and operational questions like: Where do bottlenecks occur in sampling, extraction, sequencing, and data processing? What are the limits of detection of rare species and variants? And what happens at the interface between agricultural, urban, and natural environments?”

IMSS member Laura Schnell receives the 2024 – 25 Saskatchewan Lieutenant Governor Scholarship.
Saskatchewan Lieutenant Governor Scholarship Recipients Announced
“The 2024-25 Saskatchewan Lieutenant Governor Scholarship recipient is Laura Jean Schnell, a graduate student pursuing a Doctorate of Philosophy in Microbial Genetics and Toxin Production from the University of Regina. Her research will investigate the quality of recreational, agricultural and drinking water in Saskatchewan. This research will inform water management and aim to reduce costs for livestock producers, identify potential risks to water security and provide a foundation for further water quality improvements.”
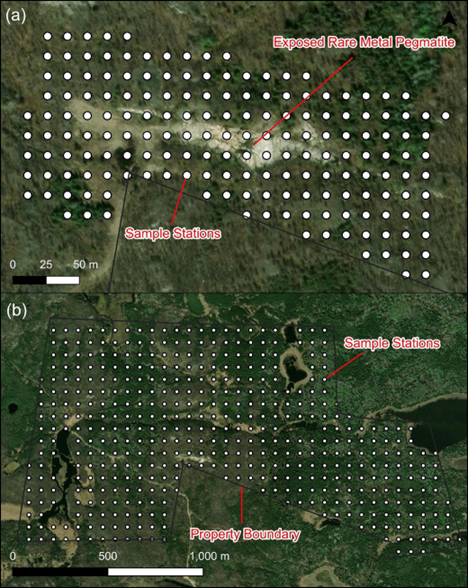
Pan American Energy Announces Research Collaboration with the University of Regina, including Investigation of Geomicrobiology for Identifying Drill Targets at the Big Mack Lithium Project – Feature in the Financial Post
“As part of the collaboration, the Company is planning a comprehensive field prospecting and sampling program utilizing eight students specializing in geosciences from the U of R at the Big Mack Lithium Project (“Big Mack” or the “Project”), including the planned collection of soil microbiology, soil chemistry, rock chemistry, ground water chemistry, soil mineralogy, environmental DNA, and vegetation biochemistry using the base layer of magnetics collected by the Company in 2023. Laboratory analyses will be conducted by Dr. Cameron and his team in the Institute for Microbial Systems and Society at the U of R as well as by the Saskatchewan Research Council, with the intent of using geomicrobiology to help generate drilling targets at Big Mack.”

“Current tests for COVID-19 and other infectious diseases are often based on polymerase chain reaction (PCR), a common technique used in labs to detect the DNA or RNA from viruses or bacteria that cause disease,” explains Cameron. “The technology is fast and efficient for detecting pathogens we already know about. However, PCR is unable to detect new or unknown pathogens that aren’t yet well-characterized. A further complication is that known pathogens can escape PCR detection by evolving new genetic sequences. The most important information for fighting infectious disease is identifying the causative agent — but, you can’t detect what you don’t test for.
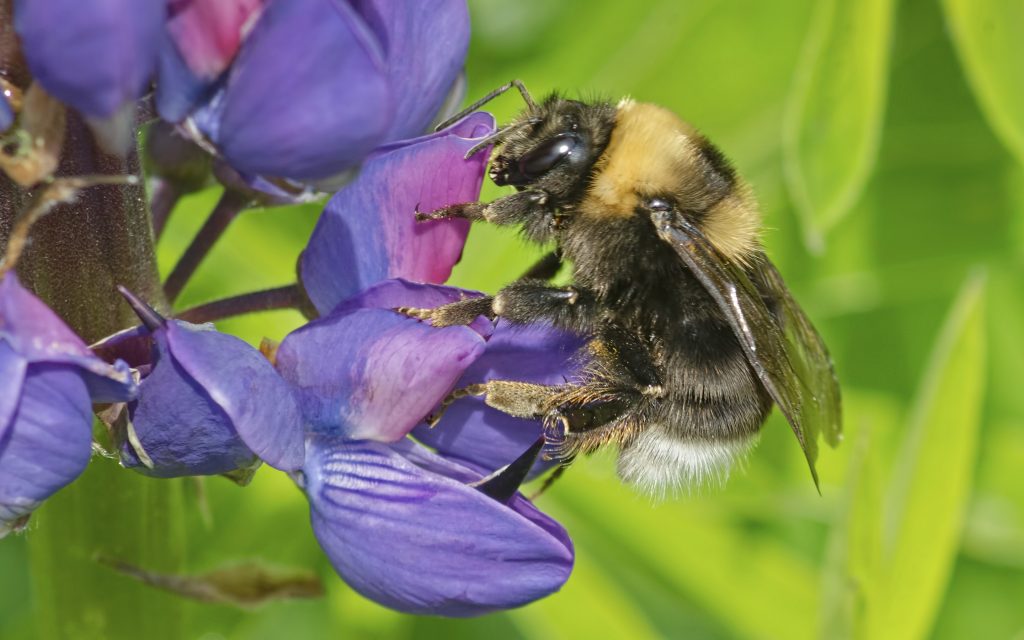
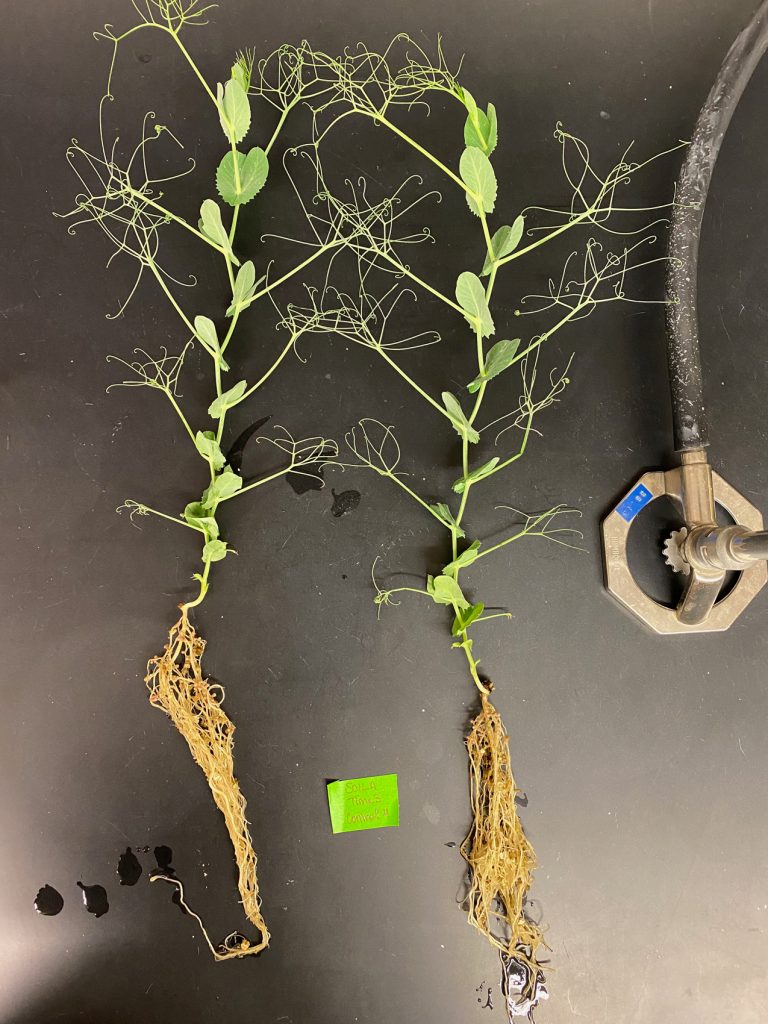


“Weeds, bees, and mould” – feature in Discourse Magazine
“It’s imperative that we figure out why our bees are disappearing. They positively impact entire ecosystems and they’re important to farmers…My preliminary research shows a potential new fungal pathogen closely related to yeast in at-risk bee species.” -About Kirsten Palmier’s research.
International posting: https://bienen-nachrichten.de/2020/neue-pathogene-bei-hummeln/730
“Aphanomyces, which kills both peas and lentils at their roots, is difficult to get rid of because it can live in soil for up to 10 years…The soil’s microbiome-which is all the bacteria and the fungus that grow around the roots-may prove to be beneficial and stop the infection.” -About Nikki Burnett’s research.
“Innovative inoculants” – feature in Discourse Magazine
https://www.discoursemagazine.ca/innovative-inoculants/2019/04/18/
“Chris Yost, biology professor and co-director of the Institute for Microbial Systems and Society, and his research team of postdoctoral fellows recently won the Award of Innovation at the Regina Chamber of Commerce 2019 Paragon Awards for their work with Lallemand Inc., a business that develops and commercializes microbe-based technologies.”
“Battling bacteria” – feature in Discourse Magazine
https://www.discoursemagazine.ca/wp-content/uploads/2019/04/discourse-fw16-17.pdf
“Research focused on understanding antibiotic resistance and developing diagnostic tools to deal with it is a priority in Caeron’s group within the Integrated Microbial Systems and Society (IMSS) Research Laboratory. They have had success on both fronts and have already had the satisfaction of seeing what they are learning put into practice combatting antibiotic resistance in Saskatchewan and elsewhere.”
“U of R researchers using genetics to detect and control infectious disease” – Regina Leader Post Article
“…Our lab is looking at how [antibiotic resistant] genes are used by the organism — when does it turn them off and on,” Cameron said. “What we’re finding is that even some of the most resistant organisms turn off their resistance sometimes. The more we can understand what induces them to turn off the resistance, then we can hit them when they make themselves vulnerable or we can trick them into turning it off.”
Funding Partners